Scientists at EPFL have developed a technique that can be a game-changer for genetics by making the characterization of DNA-binding proteins much faster, more accurate, and efficient.
Genes hold the DNA code for producing all the proteins of the cell. To begin this process, genes require a huge family of DNA-binding proteins called transcription factors, which are of enormous interest to biologists today. However, we still know very little about many transcription factors because of their large number, their ability to combine into different pairs, and the difficulty of studying their DNA-binding properties in the lab. Now, EPFL scientists have developed a microfluidics-based technique that can greatly speed up the process, with only a minimum of materials. Publishing in Nature Methods, the researchers have already used it to determine the DNA-binding properties of over 60 transcription factors, including nine new ones.
Transcription factors
Mammals — including humans — have between 1300-2000 transcription factors, many of which combine with others into “heterodimers” in order to bind genes and induce their transcription into RNA. Heterodimers are estimated to range between 3000 and 25,000.
Consequently, the number of possible combinations can be very high. But we also need to understand their DNA-binding properties, e.g. their affinity and specificity for DNA. This is key if we are to ever exploit transcription factors for biotechnological or pharmaceutical purposes in the future. But profiling transcription factors is a daunting and very complex task, since it requires relatively large amounts of hard-to-make transcription factors.
There are several databases available, but in total they cover only around 500 single transcription factors, and only a fraction of heterodimers. Progress in this field is slow, and the available profiles in these databases are not very good in predicting which genes these transcription factors work on.
A microfluidics approach
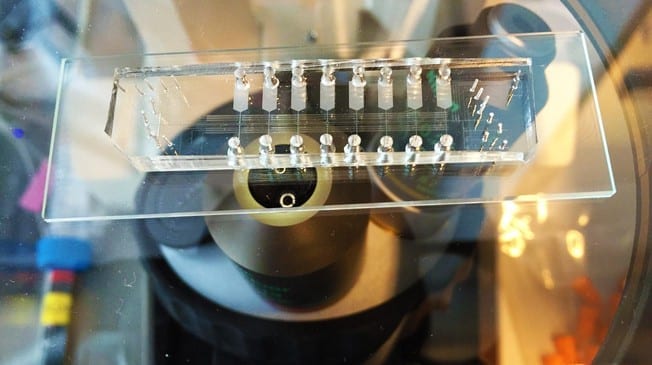
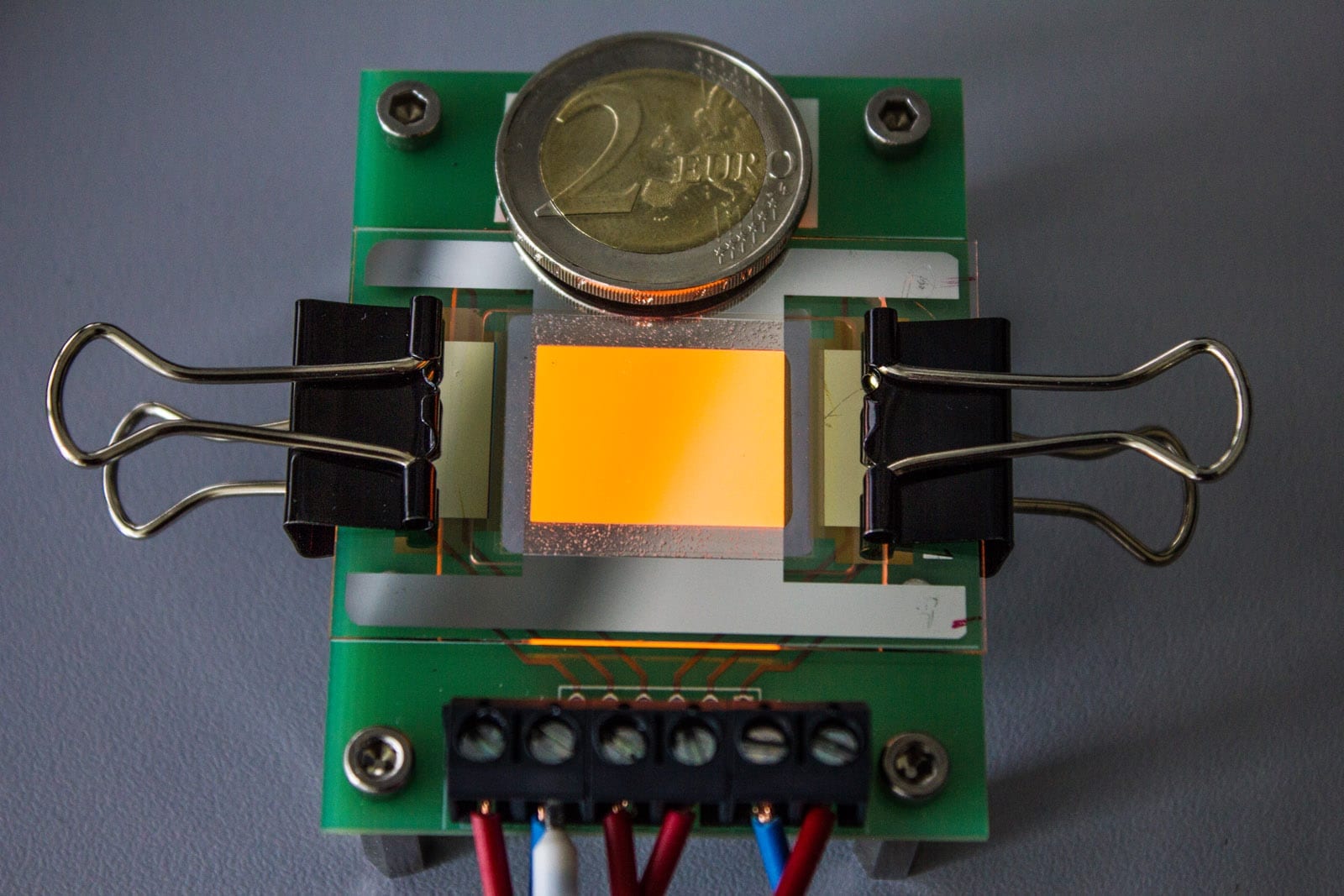
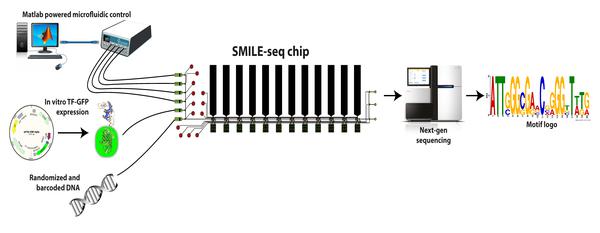
The lab of Bart Deplancke at EPFL’s Institute of Bioengineering has now invented a new technique called SMiLE-seq, which can greatly speed up the process with only tiny amounts of transcription factors. The technique uses microfluidics, which control tiny amounts of liquids in equally tiny spaces. Microfluidics is fast-becoming an area of excellence at EPFL, bringing together a number of different fields and disciplines.
SMiLE-seq works by attaching small amounts of the transcription factor (or factors when studying heterodimers) in a microfluidic device — this is a chip with micrometer-size channels that allow liquid to flow through. Once the transcription factors attach to the chip’s surface, a large library of random DNA is pumped into the chip and flows over them. This allows the transcription factors to recognize their corresponding DNA sequences. The transcription factor-DNA complex is then physically trapped, while the DNA that is not bound is washed away.
Next, the bound DNA is taken off the device and prepared for sequencing to identify which part of it got caught by the transcription factors. This information is fed into specialized software that works out the DNA-binding properties of the transcription factors or heterodimers. In turn, this helps to better predict their DNA-binding profiles in living cells.
SMiLE-seq offers three advantages: First, it cuts down on the amount of transcription factors, as it only needs picograms of them. Second, it speeds up the process from days to less than an hour. And finally, SMiLE-seq is not limited by neither the length of the DNA target sequence, nor is it biased toward stronger affinity protein-DNA interactions.
Deplancke’s team used SMiLE-seq on 67 full-length human, mouse, and Drosophila transcription factors, successfully analyzing several that have never been studied before. He next plans to exploit the technique’s versatility for other molecules, e.g. RNA. His team has filed a patent, and a startup will take the concept of SMiLE-seq into the commercial world.
Learn more: Using microfluidics to improve genetics research
[osd_subscribe categories=’dna-technology’ placeholder=’Email Address’ button_text=’Subscribe Now for any new posts on the topic “DNA TECHNOLOGY”‘]
Receive an email update when we add a new DNA TECHNOLOGY article.
The Latest on: DNA technology
[google_news title=”” keyword=”DNA technology” num_posts=”10″ blurb_length=”0″ show_thumb=”left”]
via Google News
The Latest on: DNA technology
- Non-Coding DNA Could Explain Childhood Cancer’s Radiotherapy Resistanceon May 2, 2024 at 1:48 am
Scientists at St. Jude Children’s Research Hospital have discovered non-coding genetic variants contributing to chemotherapy resistance in acute lymphoblastic leukemia.
- LPD solves the 1976 murder case of Elizabeth Ann Price with help of new DNA technologyon May 1, 2024 at 10:03 am
In 2002, evidence was submitted to the Lubbock DPS Lab four separate times, hoping that new DNA technology would break the case. However, it did not. The Texas Rangers received the SAKI grant in March ...
- Lubbock Police solves decades old homicide through DNA testingon May 1, 2024 at 8:13 am
The Metropolitan Special Crimes Unit, with the help of multiple law enforcement agencies and DNA testing, has solved the 48-year-old murder of Elizabeth Ann Price.After nearly five decades, it is now ...
- Here’s why Cathie Wood’s Ginkgo Bioworks (DNA) stock has divedon April 30, 2024 at 3:13 am
Cathie Wood, the founder of Ark Invest, is under intense pressure as most of her ETFs implode. The Ark Innovation Fund (ARKK) ETF has crashed by 72% from its highest level in 2021 while the Ark ...
- Man Does DNA Test, Not Prepared For What Comes Back 'Unusually High'on April 30, 2024 at 2:13 am
The designer's genetic profile showed an astonishing amount of Neanderthal DNA, which has captured the attention of millions online—and a scientist.
- At-home DNA test reveals man’s unusual Neanderthal connectionon April 29, 2024 at 8:36 am
Jori Waskahat did an at-home DNA test and found out that he had a Neanderthal DNA. His Instagram post went viral and sparked a much larger discussion.
- Environmental Change, Written in the DNA of Birdson April 29, 2024 at 6:59 am
Two studies of California bird populations show how shifting environments can rewrite animals’ genomes — for better or worse.
- Here is a look at one of the labs that helped uncover new DNA evidence in Lake Oconee murderon April 25, 2024 at 2:07 pm
Police don’t believe the father knew about the pregnancy, but that lead helped them identify Baby India’s biological mother, 40-year-old Karima Juani, who has now been charged with attempted murder, ...
- Vast DNA tree of life for plants revealed by global science team using 1.8 billion letters of genetic codeon April 24, 2024 at 8:00 am
A new paper published today (April 24) in the journal Nature by an international team of 279 scientists led by the Royal Botanic Gardens, Kew presents the most up-to-date understanding of the ...
via Bing News