via Stanford University
The software targets, collects, and sequences specific genes without sample preparation or extensive sequencing of surrounding genetic material
A team of Johns Hopkins University researchers has developed new software that could revolutionize how DNA is sequenced, making it far faster and less expensive to map anything from yeast genomes to cancer genes.
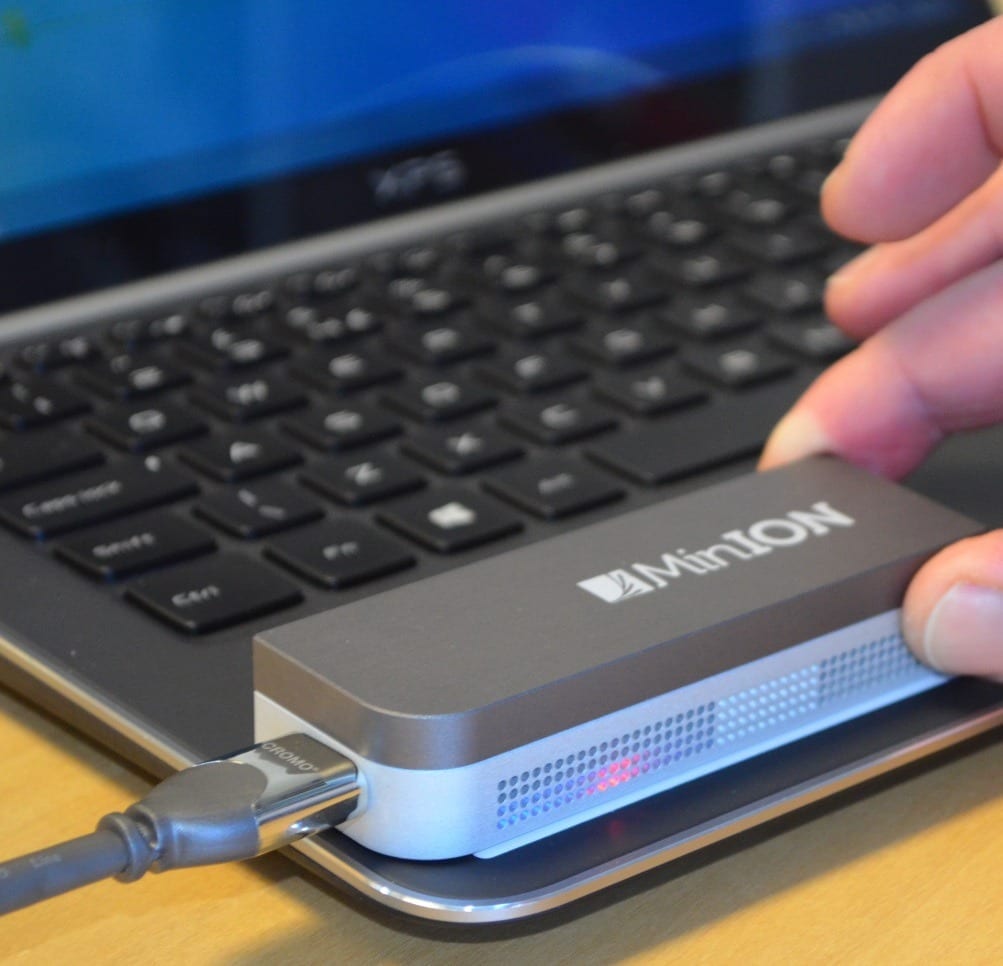
The software, detailed in a paper published in Nature Biotechnology, can be used with portable sequencing devices to accelerate the ability to conduct genetic tests and deliver diagnoses outside of labs. The new technology targets, collects, and sequences specific genes without sample preparation and without having to map surrounding genetic material like standard methods require.
“I think this will forever change how DNA sequencing is done,” said Michael C. Schatz, a Bloomberg Distinguished Associate Professor of computer science and biology and senior author of the paper.
“I THINK THIS WILL FOREVER CHANGE HOW DNA SEQUENCING IS DONE.”
Michael C. SchatzBloomberg Distinguished Associate Professor of computer science and biology
The new process shrinks the time it takes to profile gene mutations, from 15 days or more to just three. This faster timeline allows scientists to understand and diagnose conditions almost immediately, while saving time and money by eliminating preparation and additional analysis.
“In cancer genomics there are a few dozen genes known to increase cancer risk, but with a standard sequencing run, you would have to sequence the whole genome just to read off those few genes,” Schatz said, adding that adaptive sequencing allows researchers to “pick and choose which molecules we want to read and which can be skipped.”
To provide a sense of how much this invention speeds up sequencing, Schatz relates it to finding a movie on Netflix. The standard method of sequencing would require someone to watch every second of every movie on Netflix to find what they want. Instead, adaptive sequencing eliminates hours of watching irrelevant content by quickly recognizing unwanted movies and skipping to the next entry.
The open-source software’s algorithm was written by lead author Sam Kovaka, a Johns Hopkins doctoral student. Its acronymic name, UNCALLED, stands for Utility for Nanopore Current Alignment to Large Expanses of DNA.
It took two years to code, develop, and test the software, and another year to refine it enough to produce results worthy of publication, Kovaka said.
“UNCALLED allows for unprecedented flexibility in targeted sequencing,” he added. “The fact that it’s purely software-based means researchers can target any genomic region with no added cost compared to a normal sequencing run, and they can easily change targets just by running a different command.”
The process identifies DNA molecules as they pass through tiny electrified holes, or “nanopores,” inside devices called nanopore sequencers, which are cellphone-sized versions of the bulky machines used in labs. The software reads the data and checks it against a specified genome’s reference sequence within a fraction of a second. Desired molecules are allowed to pass through the pore to be fully mapped. But if an undesirable molecule is detected, the software reverses the voltage in the nanopore, physically ejecting the molecule to make room for the next.
“It’s like a nightclub doorman allowing desired guests on a list to enter while rejecting the rest with a Taser,” Schatz explains.
The research team performed two demonstrations of UNCALLED.
The first showed that the software was able to enhance the sequencing of 148 genes known to increase cancer risk by quickly and accurately profiling all of their variants with just a single run through a portable sequencer. The software made it possible to catalogue in real time dozens of complex structural mutations in the cancer genes that a standard run would have missed.
Then the team demonstrated how the software could selectively sequence certain species collected from an environment, such as microbes living on skin or those in pond water. By rejecting molecules from known microbes (such as E. coli), the software was able to efficiently sequence the remaining molecules, which revealed a less-understood yeast genome.
UNCALLED can operate on standard hardware used for nanopore sequencing without requiring special reagents or accelerators. The selection of genes or genomes to sequence is controlled entirely in the software and can be changed at any time.
The Latest Updates from Bing News & Google News
Go deeper with Bing News on:
DNA sequencing
- Sequence Listings in Patent Applications: USPTO Adopts Updated WIPO Standard
In particular, those examples illustrate how to properly represent a DNA molecule in the sequence listing by including two sequences (i.e., two SEQ ID NOs) within the sequence listing when the ...
- DNA Sequencing Market Size Anticipated to Soar to New Heights in the Future
DNA Sequencing Market is valued at approximately USD 4.7 billion in 2019 and is anticipated to grow with a healthy growth rate of more than 11.3% over the forecast period 2020-2027. DNA sequencing is ...
- Latest biotech cuts hit pioneering DNA sequencing company, Verily's eye-focused joint venture
Nearly 150 Bay Area life sciences jobs will be cut as DNA sequencing pioneer PacBio shaves positions at its Menlo Park headquarters and a South San Francisco joint venture between ...
- DNA sequencing reveals cause of rare neurological disorder
After a 25-year-long search, a new sequencing technique has helped researchers pinpoint the genetic cause of SCA4, a rare movement disorder. Spinocerebellar ataxia 4 (SCA4) is a rare genetic condition ...
- Nanopore DNA Sequencing: A Powerful Tool for Genetic Analysis
What is Nanopore DNA Sequencing? Nanopore DNA sequencing is a cutting-edge technology that enables the direct, real-time analysis of long DNA or RNA fragments. Unlike conventional sequencing methods ...
Go deeper with Google Headlines on:
DNA sequencing
[google_news title=”” keyword=”DNA sequencing” num_posts=”5″ blurb_length=”0″ show_thumb=”left”]
Go deeper with Bing News on:
Utility for Nanopore Current Alignment to Large Expanses of DNA
- What are today's home equity loan and HELOC interest rates?
The following rates are current as of May 10, 2024, according to Bankrate's home equity loan and HELOC averages. Note that these are nationwide rates. Average rates vary state by state ...
- New Jersey American Water Ranks #1 in J.D. Power 2024 Water Utility Residential Customer Satisfaction Study in Northeast Large Region
CAMDEN, N.J.--(BUSINESS WIRE)--New Jersey American Water has received the J.D. Power award for ranking highest in customer satisfaction among large water ... 2024 U.S. Water Utility Residential ...
- Enhanced CRISPR method enables stable insertion of large genes into the DNA of higher plants
Scientists at the Leibniz Institute of Plant Biochemistry (IPB) have succeeded for the first time in stably and precisely inserting large gene segments into the DNA of higher plants very efficiently.
- Resolving Large DNA Fragments with Ease
Because these insertions are large, scientists perform their quality control assessments using pulsed field gel electrophoresis (PFGE), which resolves big DNA fragments. However, traditional PFGE ...
- Large-Scale DNA Study Reveals Where People From India Originally Come From
Airspace shut down and major shipping lane closed after huge failed missile launch People Who Were Adults In The '80s Are Sharing What Pop Culture Tends To Leave Out About The Decade The 10 ...
Go deeper with Google Headlines on:
Utility for Nanopore Current Alignment to Large Expanses of DNA
[google_news title=”” keyword=”Utility for Nanopore Current Alignment to Large Expanses of DNA” num_posts=”5″ blurb_length=”0″ show_thumb=”left”]