Whole genome sequencing of tumour cells could help predict the prognosis of a patient’s cancer and offer clues to identify the most effective treatment, suggests an international study published today in Nature Medicine.
Previously, it was like going on a voyage with only a limited map, but now, with whole genome sequencing we have a much better, more detailed map and know the best route to reach our destination
Serena Nik-Zainal
Our DNA, the human genome, comprises of a string of molecules known as nucleotides. These are represented by the letters A, C, G and T. Sometimes, changes occur in the ‘spelling’ of our DNA – an A becomes a G, for example. These changes, known as mutations, can be caused by a number of factors, some spontaneous, others environmental, such as exposure to tobacco smoke or to ultraviolet light, and all leave characteristic signatures on the genome.
As cells divide and multiply, they make copies of their DNA, so any spelling mistakes will be reproduced. Over time, the number of errors accumulates, leading to uncontrolled cell growth – the development of tumours.
Whole genome sequencing (WGS) is a technique that involves reading the entire genetic blueprint of a cancer cell and comparing it to a patient’s healthy cells to see how the DNA has mutated. By studying all the mutations present in a whole cancer genome and seeking all the signatures in them, it is possible to identify the various factors that have acted upon the tumour.
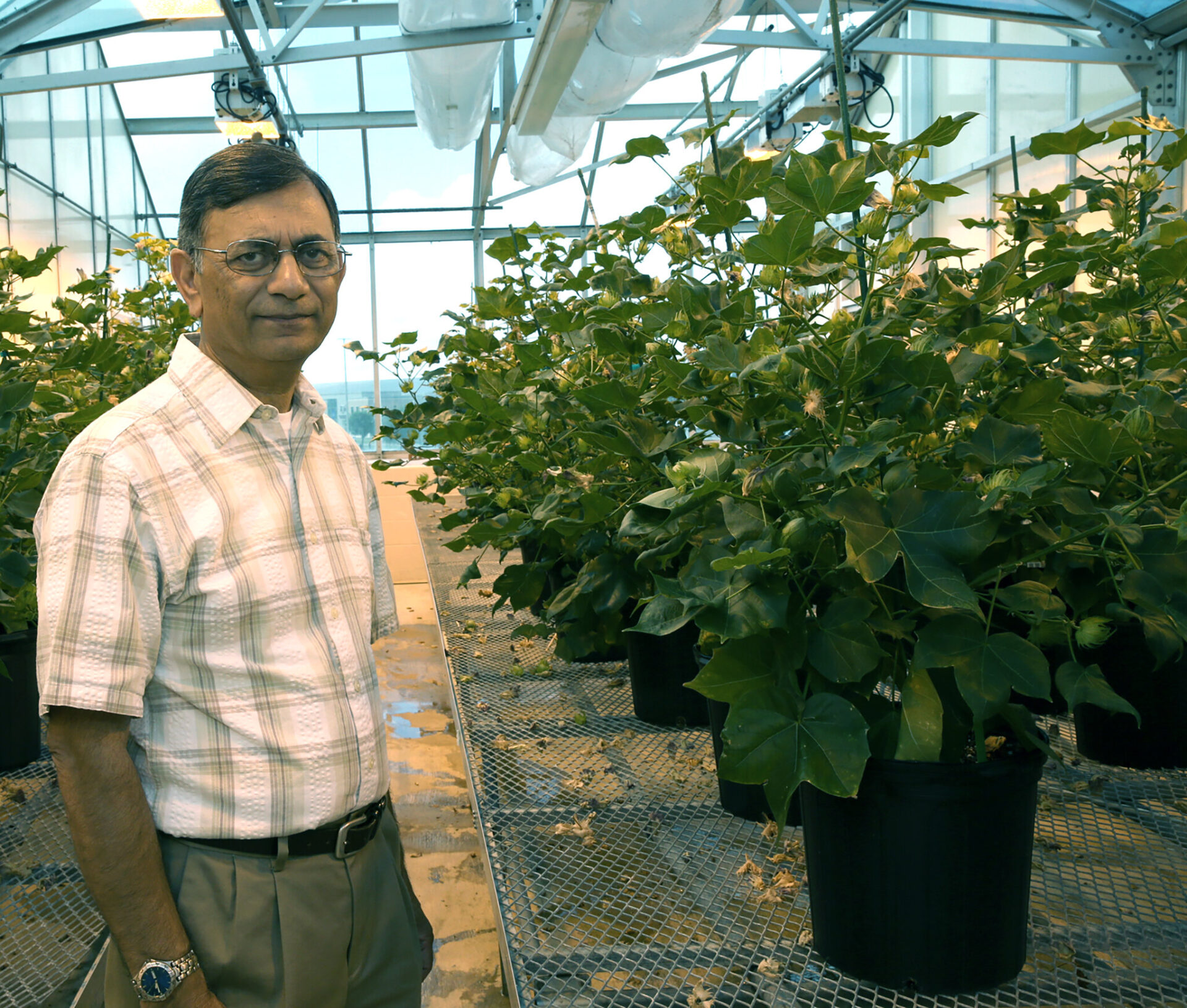
To understand whether WGS might be useful in a clinical setting, Cambridge researchers teamed up with colleagues in Sweden running a population-wide project called SCAN-B, which has been recruiting all women diagnosed with breast cancer in the South of Sweden since 2010. This was critical as SCAN-B has a large amount of clinical outcome data.
This international collaboration of researchers used WGS to analyse tumours from patients who had been diagnosed as having triple negative breast cancers. These cancers are so-called because they lack three key molecules known as receptors. They account for around 9% of breast cancers and are associated with poorer outcomes. They are also more common amongst women with African and Asian ancestry.
“Whole genome sequencing gives us a complete view of the cancer genome. It reveals many things that we couldn’t see previously, because we simply did not look for them,” explains Dr Serena Nik-Zainal from the Medical Research Council Cancer Unit at the University of Cambridge, who led the study.
“Having a complete cancer genome map for each patient helps us to understand what has caused each patient’s tumour and treat each individual more effectively. Previously, it was like going on a voyage with only a limited map, but now, with whole genome sequencing we have a much better, more detailed map and know the best route to reach our destination.”
The researchers then applied a machine learning algorithm called HRDetect, which they had previously developed to identify tumours with signatures caused by mutations in the BRCA1 or BRCA2 genes. Having a variant of either of these two genes greatly increases an individual’s risk of developing breast cancer and a relatively new class of anti-cancer drug called PARP-inhibitors have been developed specifically for these tumours. HRDetect scores had previously suggested that a greater proportion of women in the general population could have tumours that are very similar to BRCA1/BRCA2-mutant cancers.
Taking the scores, the team categorised each patient as either high, intermediate or low scoring.
Patients who scored highly were those that had the best outcomes using current treatments for triple negative breast cancers – they are also most likely to respond to PARP inhibitors.
Surprisingly, those with intermediate scores had the poorest outcomes. Current triple negative breast cancer treatments had limited effectiveness suggesting that new approaches would be necessary to tackle these cancers. However, the genetic changes and signatures revealed through WGS gave clues to the mechanisms driving these tumours, which in turn may help inform the development of new drugs.
Those patients with low scores also have poor outcomes, though not as badly as those in the intermediate group. However, the WGS profile in some of these tumours suggested biological abnormalities that could potentially be targeted by existing drugs or drugs currently going through clinical trials, such as so-called checkpoint inhibitors or AKT inhibitors.
“Using whole genome sequencing, we can truly discriminate tumours that may or may not respond to current drugs among triple negative breast cancer patients, a type of breast cancer that we still struggle to treat well,” says first author Dr Johan Staaf from the Department of Clinical Sciences, Lund University, Sweden.
“But importantly, this approach also gives us clues to some of the mechanisms that are going wrong in the poor-outcome tumours, and hence how we might treat those tumours differently or how we might develop new drugs.”
The speed of sequencing technology has progressed such that WGS can be carried out in 24 hours, with another 24-48 hours for analysis of the data. In theory, therefore, it should be possible to offer whole genome screening as a matter of course to all patients, allowing a personal readout of their tumour and potential treatment options.
“The potential for whole genome sequencing to open up a personalised approach to treating cancer is huge,” says Dr Nik-Zainal. “In the past, the cost and issues with managing the huge volume of data created barriers to its widespread application. But we are moving closer to a time where it can be routinely offered to all patients, with the potential to transform the management of even difficult-to-treat cancers.”
Learn more: Study highlights potential of whole genome sequencing to enable personalised cancer treatment
The Latest on: Whole genome sequencing
[google_news title=”” keyword=”whole genome sequencing” num_posts=”10″ blurb_length=”0″ show_thumb=”left”]
via Google News
The Latest on: Whole genome sequencing
- Why genomic sequencing of the dengue virus can help control the diseaseon May 17, 2024 at 9:00 am
As per the World Health Organization, about half of the world's population is now at risk of dengue with an estimated 100-400 million infections occuring each year.
- UP PGC empowers educators to teach DNA sequencing & other molecular biology techniqueson May 16, 2024 at 6:01 pm
During the COVID-19 pandemic, the efforts made by the University of the Philippines (UP) to develop expertise in Next Generation Sequencing (NGS) and Genomics were integral in the country’s national ...
- ProPhase Labs: Hidden Gem With An AI Kickeron May 15, 2024 at 7:43 am
ProPhase's Project ZenQ-AI aims to use AI to identify potential antibody drug candidates, with the potential for significant revenue growth. Click here to read.
- When good bacteria go bad: New links between bacteremia and probiotic useon May 14, 2024 at 12:50 pm
Researchers discovered a concerning association between bacteremia and probiotic use, particularly with Clostridium butyricum (C. butyricum) MIYAIRI 588. Whole-genome sequencing revealed that all C.
- Analysis suggests people with more copies of ribosomal DNA have higher risks of developing diseaseon May 14, 2024 at 8:00 am
Ribosomal DNA (rDNA) is present in hundreds of copies in the genome, but has not previously been part of genetic analyses. A new study of 500,000 individuals indicates that people who have more copies ...
- The Largest Whole-genome Sequencing Study in Canceron April 16, 2024 at 5:01 pm
it's very likely that everyone will move towards whole-genome sequencing because it will be cost-effective. And what that will allow us to do is to detect unusual features in people's cancers that go ...
- Whole-exome Sequencing: Opportunities in Pediatric Endocrinologyon April 14, 2024 at 5:00 pm
There are three basic paradigms in which this technology is usually applied: whole-genome sequencing (WGS), whole-exome sequencing (WES) and targeted gene sequencing (TGS). [18–19] In WGS ...
- Discovery of genetic variants linked to Alzheimer’s offers hope for cureon March 21, 2024 at 7:28 am
The breakthrough, discovered using whole genome sequencing, identified 17 significant variants associated with Alzheimer’s disease in five genomic regions. Scientists say the discovery could ...
- Whole genome sequencing identifies causal variants in CMTon December 14, 2023 at 6:28 am
The promise of affordable individual human genome sequencing information has nurtured a whole new industry that races to deliver the $1,000 genome. Sequencing output has increased 5–10-fold per ...
- Artikel-artikel mengenai Whole-genome sequencingon March 30, 2022 at 5:00 pm
Some argue that a controversial tool called whole genome sequencing may help find the cause. Genome sequencing is transforming the way we diagnose disease. But lack of diversity in genomic data ...
via Bing News