When you get sick, you want the right treatment fast. But certain infectious microbes are experts at evading the very anti-bacterial drugs designed to fight them.
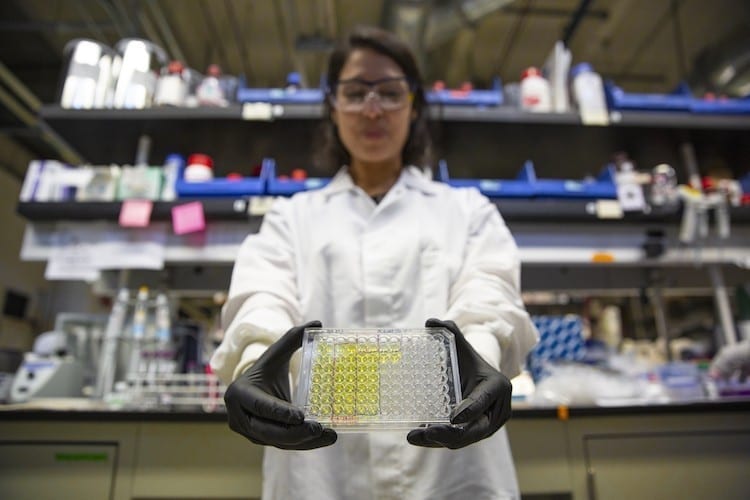
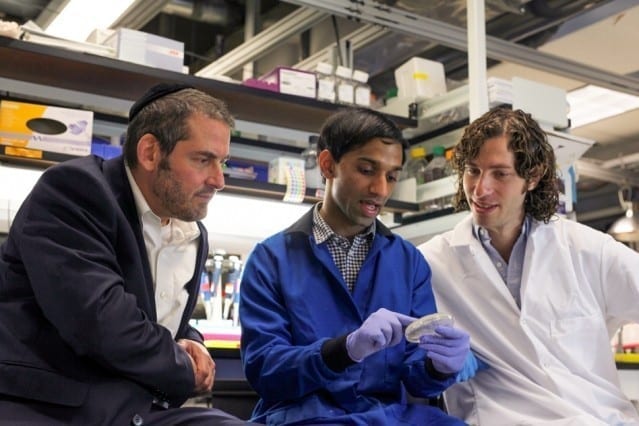
A simple and inexpensive new test developed by UC Berkeley researchers can identify bacteria that are resistant to certain classes of antibiotics in a matter of minutes. The technique could help doctors prescribe the right antibiotics for each infection, and could help limit the spread of antibiotic-resistant “superbugs,” which kill as many as 700,000 people worldwide each year.
“Health organizations around the world are supporting the development of tools that specifically identify pathogens that are resistant to antibiotics because there are limited tests available that can do it quickly,” said Tara deBoer, a postdoctoral fellow in the College of Engineering at UC Berkeley. “Our test is simple and gives us information on a short timescale.”
The test, dubbed DETECT, spots the molecular signatures of antibiotic-resistant bacteria directly in urine samples of patients with urinary tract infections. Unlike other techniques that are currently on the market, DETECT does not require expensive instrumentation and is simple enough to be applied in a point-of-care setting.
“In theory, DETECT will allow you to diagnose antibiotic-resistant urinary tract infections in a doctor’s office just by collecting urine and mixing it with the DETECT reagents,” said Niren Murthy, a professor of engineering at Berkeley.
The team is working to perfect the technique so that it can also be used to detect antibiotic resistant bacteria in blood samples.
“Drug-resistant infections are a silent pandemic that actually kill more people every year than Zika or Ebola,” said Lee Riley, professor of epidemiology and infectious diseases in the School of Public Health at UC Berkeley. “The faster you can start the right drug, the better the chances of survival or avoiding complications.”
The study, which was conducted as part of the Consortium for Research on Antimicrobial Resistant Bacteria (CRARB), which includes Berkeley researchers in the College of Engineering and the School of Public Health, appears on the Oct. 18 cover of the journal ChemBioChem.
Chopping up antibiotics
Many common early-generation antibiotics, including penicillin, amoxicillin and ampicillin, are based around a molecular structure called beta-lactam, which blocks bacteria from building cell walls, making it impossible for microbes to grow and reproduce.
However, as use of these antibiotics has soared over the past 80 years, certain infectious bacteria, including strains of E. coli, Salmonella and Shigella, have evolved to produce enzymes called beta-lactamases that chop up these antibiotics and render them useless.
DETECT works by identifying the presence of beta-lactamases in urine samples. “What our technology does is detect the molecules that are actually breaking down the antibiotics,” deBoer said.
While the basic technique for detecting beta-lactamases has already been developed, it is not sensitive enough to spot the relatively small concentrations of beta-lactamases in patient urine samples. For this technique to work, bacteria from a patient sample must first be cultured in a lab, which can take two to three days — long enough for a simple bacterial infection like a urinary tract infection to invade the kidneys or the blood.
The DETECT technique uses an enzymatic chain reaction to boost the signal from beta-lactamases by a factor of 40,000, high enough to allow detection of the presence of these enzymes in urine samples. With DETECT, a patient who tests positive for a urinary tract infection that is resistant to early-generation antibiotics can immediately be treated with a more powerful antibiotic or alternative agent.
The team tested DETECT on 40 urine samples collected from patients suspected of having a urinary tract infection, and found that approximately one-quarter of them had antibiotic-resistant infections.
“DETECT tells you not only who has antibiotic-resistant infections but also tells you who could be treated by early-generation antibiotics, allowing you to spare higher-end antibiotics and slow the spread of drug resistance,” Murthy said.
From the lab to the doctor’s office
DeBoer is now collaborating with doctors and clinical lab specialists in hospitals to design easy-to-use DETECT-based devices catering to specific medical settings.
“Everybody has different needs in the hospital,” deBoer said. “Right now we have a lot of designs, but what we are doing is allowing the intended use to define what the design is going to look like.”
For example, diagnostic tools that work well in an out-patient clinic may not be as convenient for doctors working in an emergency department, deBoer said.
With the help of UC Berkeley’s start-up incubator CITRIS Foundry, deBoer has co-founded a company, BioAmp Diagnostics, which is working to commercialize the technology into a rapid diagnostic device.
The team is continuing to perfect its enzyme signal-amplification technique in hopes of soon being able to apply it to detect specific strains of bacteria as well as bacteria in the blood.
“I think we are on the verge of having this applicable in a hospital setting,” Riley said.
Learn more: New test rapidly identifies antibiotic-resistant ‘superbugs’
The Latest on: Antibiotic resistance
[google_news title=”” keyword=”antibiotic resistance” num_posts=”10″ blurb_length=”0″ show_thumb=”left”]
via Google News
The Latest on: Antibiotic resistance
- High levels of resistant bacteria found in uncooked meats and raw dog food: ‘Red flag’on May 1, 2024 at 7:09 pm
Eighty-one percent of the meat samples and 87% of the dog food samples were found to contain E. coli (Escherichia coli) that was resistant to antibiotics. The raw chicken had the highest levels of the ...
- The State Of: Antimicrobial Resistanceon May 1, 2024 at 2:44 am
We take a look at the AMR field, looking at the past, present and future to give you a comprehensive overview of the research landscape.
- How does CRISPR edit antimicrobial-resistant bacteria?on May 1, 2024 at 2:27 am
Research has recently been presented highlighting the role that CRISPR-Cas technology can play in tackling antimicrobial resistance.
- Raw Meat for Pets: Possible Link to Antibiotic-resistant Bacteriaon April 29, 2024 at 10:28 pm
Study shows raw meat fed to animals causes antibiotic resistance to critical antibiotics, which can also be acquired by pet owners when feeding companion animals.
- Antibiotic Alternative Produced by Gram-Positive Bacteriaon April 29, 2024 at 10:15 pm
Due to increasing antibiotic resistance in pathogens causing infections, the development of new antibacterial substances is needed. Scientists are testing out a new group of substances produced by ...
- Antimicrobial resistance crisis: 'Antibiotics are not magic bullets'on April 29, 2024 at 10:26 am
Science X is a network of high quality websites with most complete and comprehensive daily coverage of the full sweep of science, technology, and medicine news ...
- Study reveals high levels of antibiotic resistance in meat sold for consumptionon April 28, 2024 at 10:28 pm
New research presented at the ESCMID Global Congress (formerly ECCMID) in Barcelona, Spain (27-30 April) has found substantial levels of resistance to critically important antibiotics in meat sold for ...
- Antimicrobial resistance projected to kill 10 million people each yearon April 28, 2024 at 10:45 am
Dr. James Gill addresses antimicrobial resistance and its accelerating crisis, emphasizing the need for patient education.
- A vaccine to combat antibiotic resistanceon April 27, 2024 at 2:00 pm
A team of researchers at Michigan State University have outlined an approach to combating a prevalent public health issue: the development of treatment-res | Technology ...
via Bing News