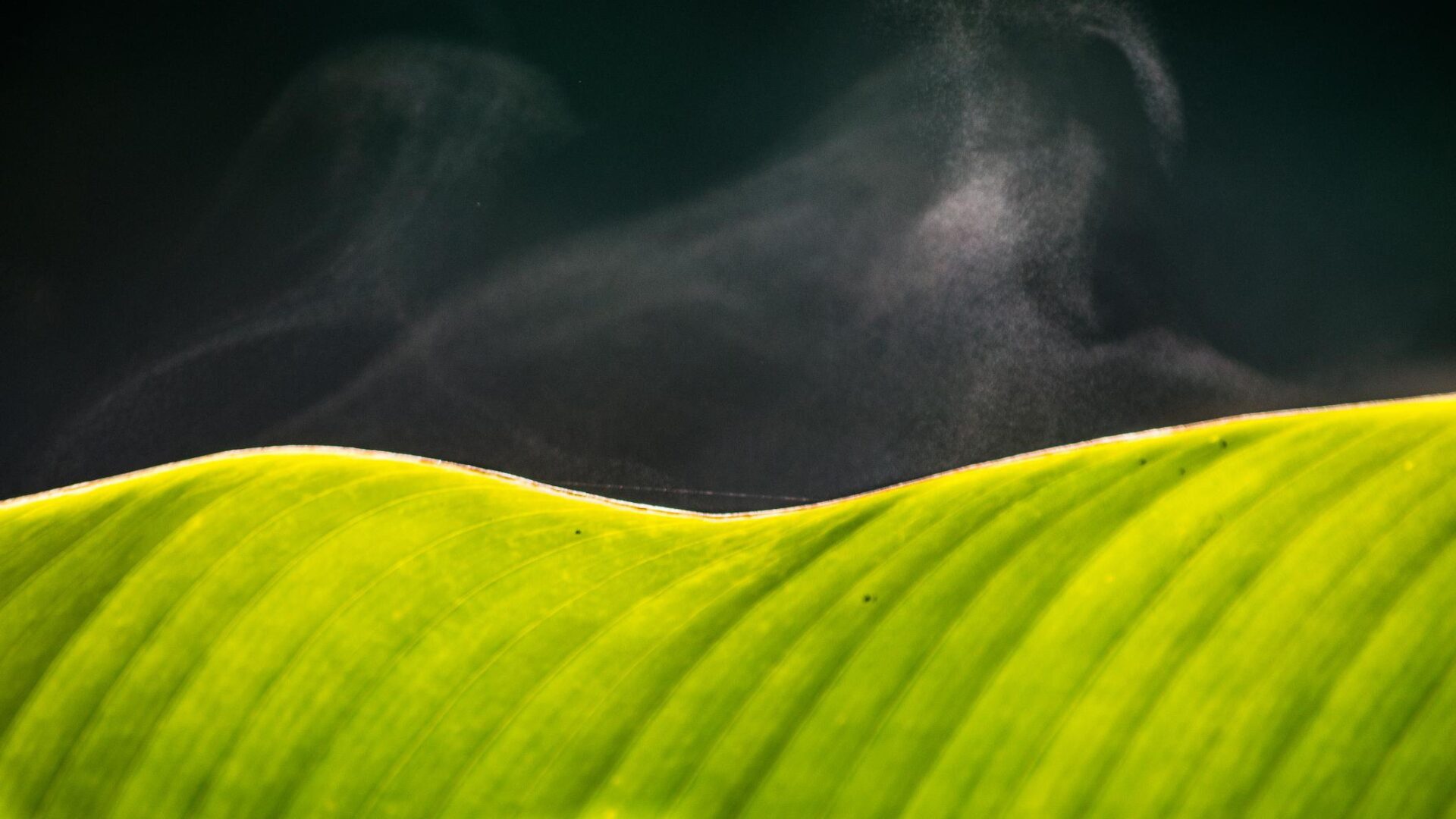
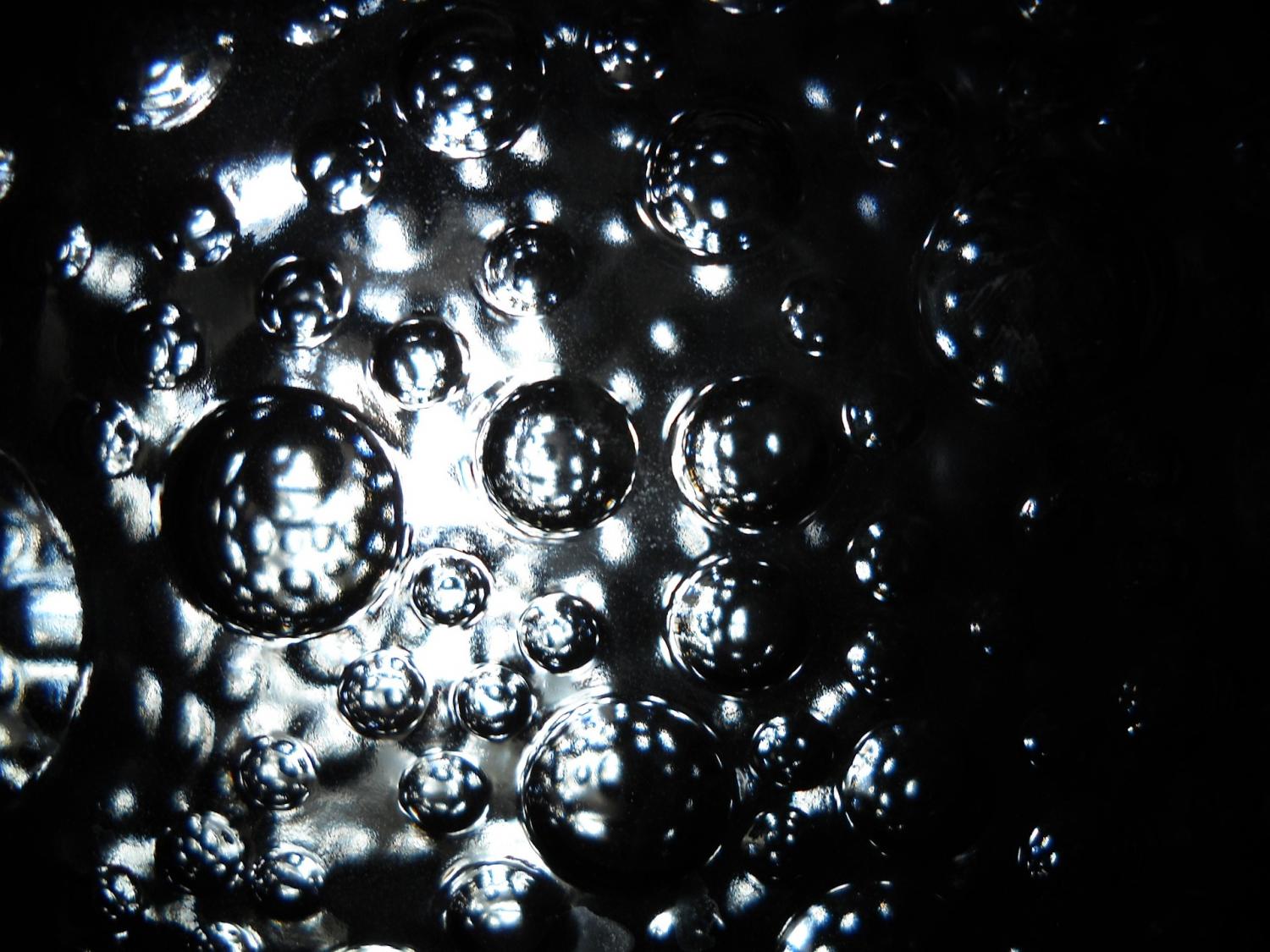
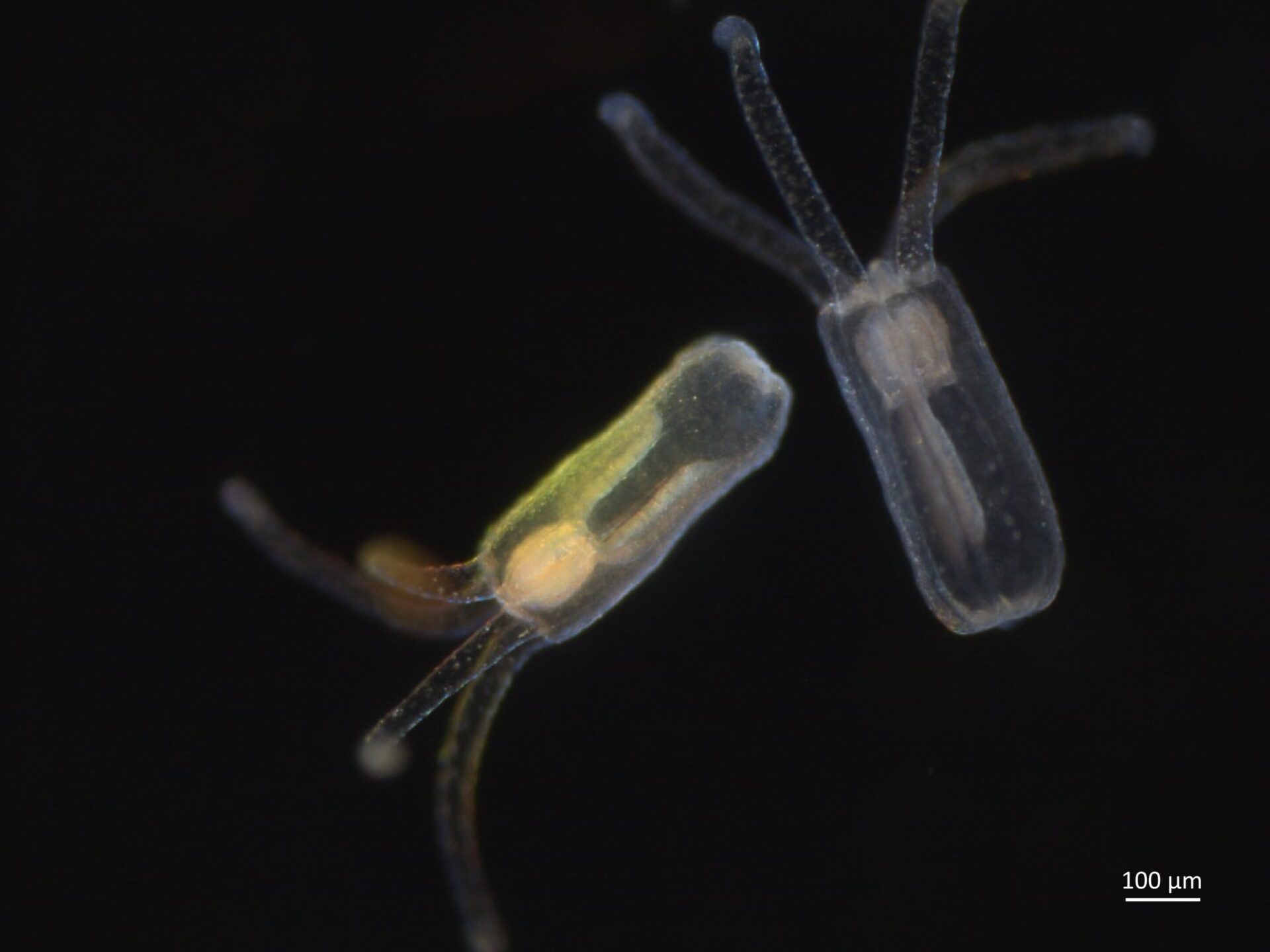
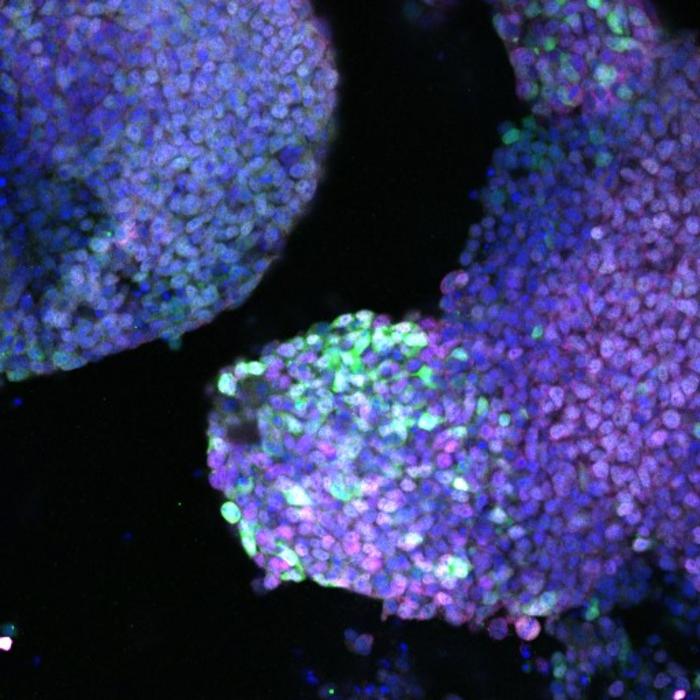
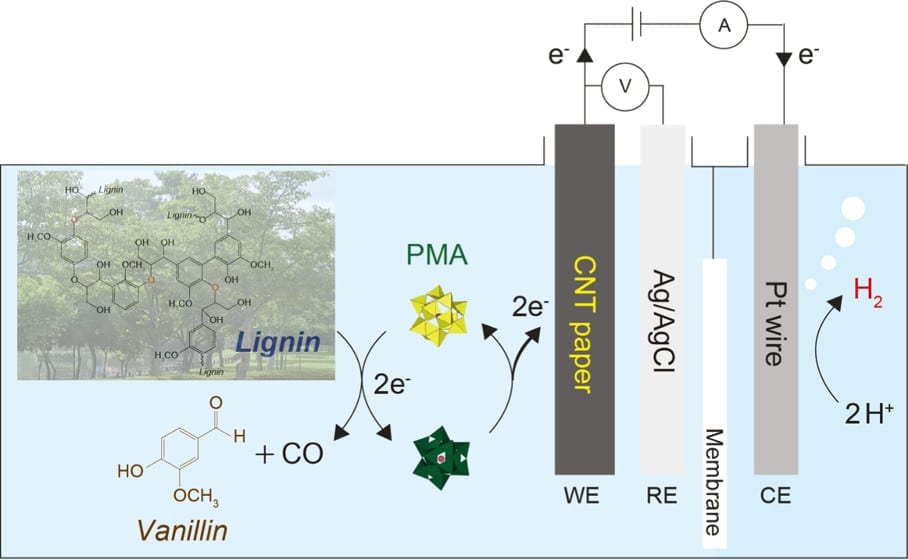
At left, a scanning electron microscope image shows the mesoporous structure of molecular-imprinted graphitic carbon nitride nanosheets. At right, a transmission electron microscope image shows the sheet’s edge and its crystalline structure. Rice researchers imprinted the nanosheets to catch and kill free-floating antibiotic resistant genes found in secondary effluent produced by wastewater plants. Courtesy of the Alvarez Research Group
Rice University material ‘traps and zaps’ floating DNA that makes bacteria resistant
It’s not enough to take antibiotic-resistant bacteria out of wastewater to eliminate the risks they pose to society. The bits they leave behind have to be destroyed as well.
Researchers at Rice University’s Brown School of Engineering have a new strategy for “trapping and zapping” antibiotic resistant genes, the pieces of bacteria that, even though theirs hosts are dead, can find their way into and boost the resistance of other bacteria.
The team led by Rice environmental engineer Pedro Alvarez is using molecular-imprinted graphitic carbon nitride nanosheets to absorb and degrade these genetic remnants in sewage system wastewater before they have the chance to invade and infect other bacteria.
The researchers targeted plasmid-encoded antibiotic-resistant genes (ARG) coding for New Delhi metallo-beta-lactamase 1 (NDM1), known to resist multiple drugs. When mixed in solution with the ARGs and exposed to ultraviolet light, the treated nanosheets proved 37 times better at destroying the genes than graphitic carbon nitride alone.
The work done under the auspices of the Rice-based Nanosystems Engineering Research Center for Nanotechnology-Enabled Water Treatment (NEWT) is detailed in the American Chemical Society journal Environmental Science and Technology.
“This study addresses a growing concern, the emergence of multidrug resistant bacteria known as superbugs,” said Alvarez, director of the NEWT Center. “They are projected to cause 10 million annual deaths by 2050.
“As an environmental engineer, I worry that some water infrastructure may harbor superbugs,” he said. “For example, a wastewater treatment plant in Tianjin that we’ve studied is a breeding ground, discharging five NDM1-positive strains for each one coming in. The aeration tank is like a luxury hotel where all bacteria grow.
“Unfortunately, some superbugs resist chlorination, and resistant bacteria that die release extracellular ARGs that get stabilized by clay in receiving environments and transform indigenous bacteria, becoming resistome reservoirs. This underscores the need for technological innovation, to prevent the discharge of extracellular ARGs.
“In this paper, we discuss a trap-and-zap strategy to destroy extracellular ARGs. Our strategy is to use molecularly imprinted coatings that enhance selectivity and minimize interference by background organic compounds.”
Molecular imprinting is like making a lock that attracts a key, not unlike natural enzymes with binding sites that only fit molecules of the right shape. For this project, graphitic carbon nitride molecules are the lock, or photocatalyst, customized to absorb and then destroy NDM1.
To make the catalyst, the researchers first coated the nanosheet edges with a polymer, methacrylic acid, and embedded guanine. “Guanine is the most readily oxidized DNA base,” Alvarez said. “The guanine is then washed with hydrochloric acid, which leaves behind its imprint. This serves as a selective adsorption site for environmental DNA (eDNA).”
Rice graduate student Danning Zhang, co-lead author of the paper, said carbon nitride was chosen for the base nanosheets because it is nonmetallic and is thus safer to use, and for its easy availability.
Alvarez noted all catalysts are efficient at removing ARGs from distilled water, but not nearly as effective in secondary effluent, a product of sewage treatment plants after solids and organic compounds are removed.
“In secondary effluent, you have reactive oxygen species scavengers and other inhibitory compounds,” Alvarez said. “This trap-and-zap strategy significantly enhances removal of the eDNA gene, clearly outperforming commercial photocatalysts.”
The researchers wrote that conventional disinfection processes used at wastewater treatment plants, including chlorination and ultraviolet radiation, are moderately effective in removing antibiotic-resistant bacteria but relatively ineffective at removing ARGs.
They hope their strategy can be adapted on an industrial scale.
Zhang said the lab has not yet run extensive tests on other ARGs. “Since guanine is a common constituent of DNA, and thus ARGs, this approach should also efficiently degrade other eARGs,” he said.
There is room to improve the current process, despite its extraordinary initial success. “We have not yet attempted to optimize the photocatalytic material or the treatment process,” Zhang said. “Our objective is to offer proof-of-concept that molecular imprinting can enhance the selectivity and efficacy of photocatalytic processes to target eARGs.”
The Latest Updates from Bing News & Google News
Go deeper with Bing News on:
Antibiotic resistant genes
- Avoid salad when on antibiotics to ‘fight bacterial resistance’
People taking antibiotics should cook food thoroughly and avoid salad vegetables to reduce antimicrobial resistance. Researchers at the University of Nottingham warned that bacteria carrying genes ...
- Mystery CRISPR unlocked: A new ally against antibiotic resistance?
CRISPR-Cas systems have revolutionized biotechnology by offering ways to edit genes like a pair of programmable scissors. In nature, bacteria use these systems to fight off deadly viruses. A recent ...
- Study Sheds Light on Preventing PXR-Associated Drug Resistance
Scientists at St. Jude Children’s Research Hospital uncovered a route to blocking activity of protein notorious for eliminating drugs, offering a potential boon to cancer therapy.
- Study finds antimicrobial resistance in soils Scotland-wide
Resistance to antibiotics has been found in the environment across Scotland, according to a new international study involving Strathclyde.
- Fears over emergence of new superbugs as antibiotic-resistant genes found in soil samples from across Scotland
Organisms with genetic defences against our most commonly used medicines are everywhere in Scotland – not good news for human and animal health ...
Go deeper with Google Headlines on:
Antibiotic resistant genes
[google_news title=”” keyword=”antibiotic resistant genes” num_posts=”5″ blurb_length=”0″ show_thumb=”left”]
Go deeper with Bing News on:
Environmental DNA
- Local researchers follow orca flukeprints in Monterey Bay
Each spring gray whales migrate north from Mexico, towing their newborn calves over the formidable Monterey Canyon for the first time. Orcas also come into the bay. to hunt the baby grays. Now local ...
- 10 unexpected ways Neanderthal DNA affects our health
For example, a 2020 study published in the journal Nature found that Neanderthal DNA on chromosome 3, which is carried by 16% of Europeans and 50% of South Asians, is associated with an increased risk ...
- Wolf or coyote? Wildlife mystery in Nevada solved with DNA testing
Nevada wildlife officials launched a huge investigation after spotting three animals believed to be wolves, which do not normally live in the state.
- AI aids analysis of lifetime environmental exposures
The idea that biology is not destiny is hardly new. Studies in twins have shown that even among identical pairs—those sharing 100% of their DNA—the same disease genes do not turn into full-blown ...
- Environmental Change, Written in the DNA of Birds
Two studies of California bird populations show how shifting environments can rewrite animals’ genomes — for better or worse.
Go deeper with Google Headlines on:
Environmental DNA
[google_news title=”” keyword=”environmental DNA” num_posts=”5″ blurb_length=”0″ show_thumb=”left”]